Amanda Hummon
Professor
414 Biomedical Research Tower
460 W 12th Ave
Columbus, OH 43210
Areas of Expertise
- Analytical
- Biochemistry
Biography
Amanda Hummon earned her A.B. in chemistry at Cornell University in 1999 with honors. She completed her graduate studies in analytical chemistry at the University of Illinois, Urbana-Champaign, in the laboratory of Prof. Jonathan V. Sweedler. Her thesis work focused on the development of mass spectrometric and bioinformatic strategies to predict and identify neuropeptides. She received her Ph.D. in 2004. Amanda participated in the annotation of the newly sequenced honey bee genome as a post-doctoral fellow in the laboratories of Prof. Gene E. Robinson and Prof. Sandra L. Rodriguez-Zas at the University of Illinois from 2004-2005. The focus of her research was constructing a methodology to utilize detected gene products to decipher an unannotated genome.
From 2005-2009, Amanda was a Sallie Rosen Kaplen Post-Doctoral Fellow at the National Cancer Institute in the laboratory of Dr. Thomas Ried. During her time in the Ried lab, she utilized RNA interference screening techniques followed by microarray analysis to elucidate genes that regulate the viability of colorectal cancer cells.
In 2009, she began her independent career as the Walther Cancer Assistant Professor in the Department of Chemistry and Biochemistry at the University of Notre Dame and was promoted to the Charles L. Huisking Associate Professor in 2015. She has been recognized with a NSF CAREER award (2014), a Society for Analytical Chemists of Pittsburgh Starter Grant Award (2011), and a Rising Star Award from the American Chemical Society (2016). Amanda moved her research program to the Department of Chemistry and Biochemistry at The Ohio State University in January 2018 and was awarded the Presidential Early Career Award for Scientists and Engineers (PECASE) in 2019. In 2020, she was awarded a Fulbright Scholar Award and will spend a semester as a Visiting Professor at the Maastricht MultiModal Molecular Imaging Institute at Maastricht University in the Netherlands.
Research Overview:
Our research interests lie at the intersection of analytical chemistry and chemical biology, with a focus on cancer biology. Cancer is a complex disease, requiring sophisticated and systematic strategies to deliver knowledge that will lead to improved treatment options for patients. Individuals trained in a wide range of scientific topics and possessing a substantial breadth of knowledge are best positioned to tackle this challenge. In the Hummon Research Group, we develop analytical methods to evaluate both the transcriptome and the proteome in cancer cells, while exploring the deregulation in cancer-associated signal transduction pathways.
Cancer arises from insults to the genome. With genomic damage, the expression levels of genes are altered from their normal state. Changes in the genome, transcriptome and proteome have been found to be highly conserved among samples from adenomas to carcinomas to metastases. Because genetic changes are commonly repeated among cancer patients, a better understanding of which genes, transcripts, and proteins are affected could have broad health implications. Therefore, the best way to understand the molecular underpinnings of cancer is to dissect the deregulated pathways that are contributing to the cancer phenotype, identify the aberrantly expressed genes and their products, and decipher their effect on downstream targets.
Biomolecule Visualization and Quantification Using Mass Spectrometry Imaging
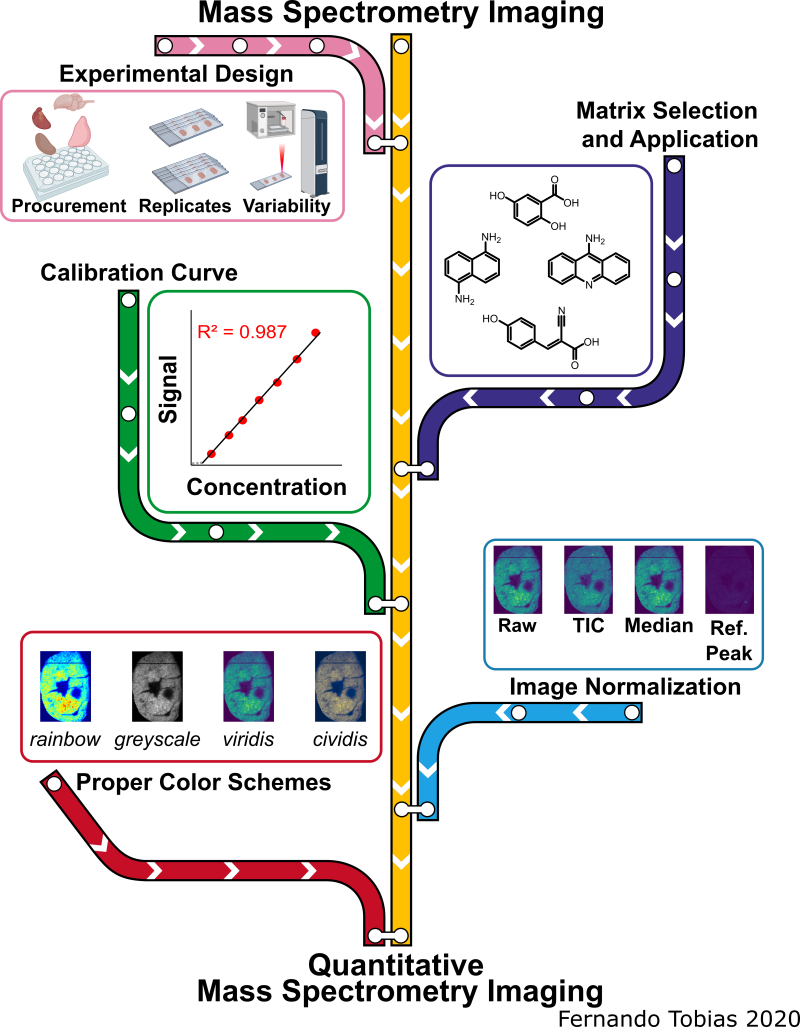
The pink line represents experimental design and sources of variability, the purple line signifies the matrix selection process and application methods, the green line represents creating calibration curves for qMSI, the blue line symbolizes the image normalization aspect, while the red line represents the selection process for the proper color scheme of imaging data. The MS images depicted in this scheme are from the same murine heart section in a different color scheme and when the signal was normalized by different methods.
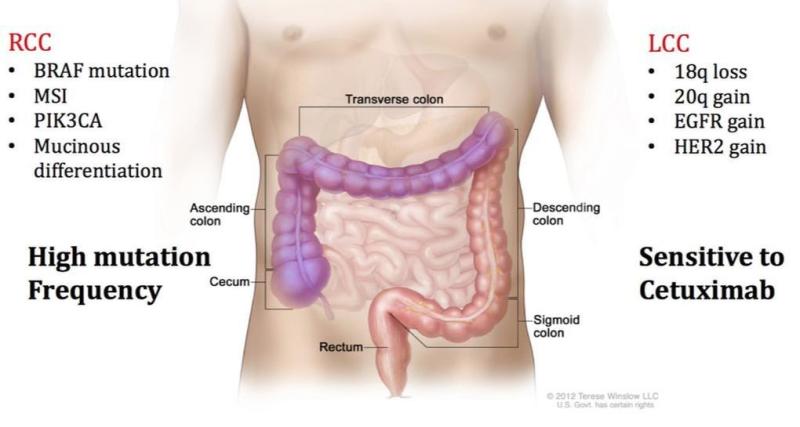
Proteomic Analysis of Right-Side and Left-Side Colon Cancer
Numerous studies have demonstrated that right-side colon cancer (RCC) and left-side colon cancer (LCC) exhibit distinct characteristics and should be treated as separate diseases. RCC spans the cecum, ascending colon, and transverse colon, while LCC includes the descending and sigmoid colon. RCC patients often have a poor prognosis compared to LCC patients because LCC patients respond more successfully to traditional FOLFOX and cetuximab treatments. RCC and LCC have distinct macroscopic and molecular characteristics. We are examining the proteomic differences found in RCC and LCC tumors to further understand the complexities of this disease.
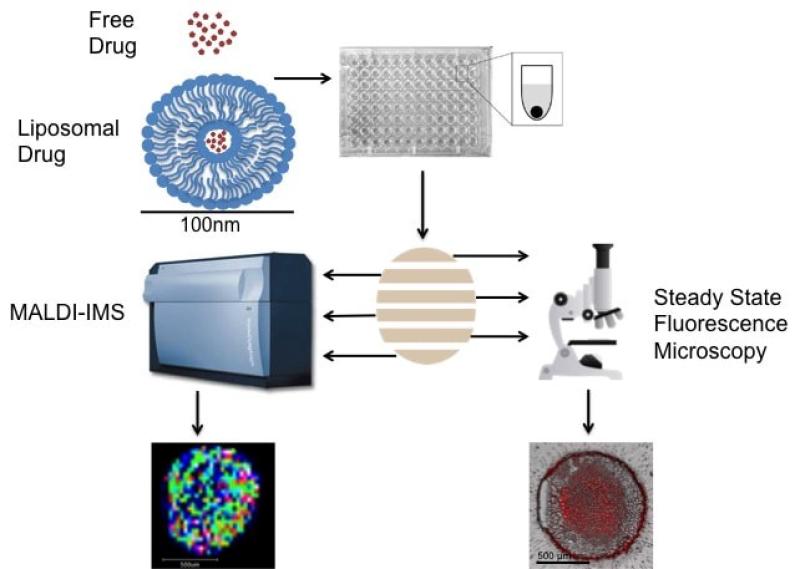
Imaging Mass Spectrometry Analysis of Liposomal Drug Delivery Systems
Cancer chemotherapeutics often fail to reach all diseased cells. To help solve this problem, researchers are investigating novel drug delivery systems. Liposomes are an attractive option due to their low toxicity, high biocompatibility, and potential to carry a large amount of a drug to the tumor site, all while avoiding being eliminated from the body. Using MALDI-IMS and fluorescence microscopy we analyze the spatial distribution of liposomal drugs in comparison to their free drug counterpart within spheroids.
Nutrient Restriction and Colorectal Cancer
The use of short-term fasting is a potential cancer treatment that could be used in tandem with current cancer regimes to increase their potency. We set out to understand the molecular mechanisms behind nutrient restriction in CRC with the use quantitative proteomics. Quantitative proteomics is a tool to survey all of the proteins in a cell and their relative abundance between biological conditions. Using this technology, we are studying the changes that occur from the addition of glucose restriction and autophagy inhibition to treatment regimens for colorectal cancer.
Recent Publications
Tobias, F and Hummon AB “Considerations for MALDI-based Quantitative Mass Spectrometry Imaging Studies” J Proteome Res. 2020 19 (9), 3620-3630 PMID: 32786684
Andrews, WT; Donahue, D; Holmes, A; Balsara, R; Castellino, FJ “In situ metabolite and lipid analysis of GluN2D-/- and wild-type mice after ischemic stroke using MALDI MSI” Anal Bioanal Chem. 2020 Feb 28. PMID: 32107573
Tobias, F, McIntosh, JC, LaBonia, GJ, Boyce, MW, Lockett, MR, and Hummon, AB “Developing a Drug Screening Platform: MALDI-Mass Spectrometry Imaging of Paper-Based Cultures” Anal Chem. 2019, 91(24): 15370–15376 PMID: 31755703
Lukowski, JK and Hummon AB “Quantitative evaluation of liposomal doxorubicin and its metabolites in spheroids.” Anal Bioanal Chem. 2019 411(27): 7087-7094. PMID: 31471684
Liu, X; Lukowski, JK; Flinders, C; Mumenthaler, S; Hummon, AB “MALDI-MSI of Immunotherapy: Mapping the EGFR-Targeting Antibody Cetuximab in 3D Colon Cancer Cell Cultures” Anal Chem. 2018 90(24): 14156-64. PMID: 30479121
Ludwig KR, Schroll MM, Hummon AB. “Comparison of In-Solution, FASP, and S-Trap Based Digestion Methods for Bottom-Up Proteomic Studies.” J Proteome Res.2018 Jul 6;17(7):2480-2490
LaBonia GJ, Ludwig KR, Mousseau CB, Hummon AB. “iTRAQ Quantitative Proteomic Profiling and MALDI-MSI of Colon Cancer Spheroids Treated with Combination Chemotherapies in a 3D Printed Fluidic Device.” Anal Chem. 2018 Jan 16;90(2):1423-1430
Liu X, Flinders C, Mumenthaler SM, Hummon AB. “MALDI Mass Spectrometry Imaging for Evaluation of Therapeutics in Colorectal Tumor Organoids.” J Am Soc Mass Spectrom. 2018 Mar;29(3):516-526
